Enzyme And Oligo Sequence Chart
ADVERTISEMENT
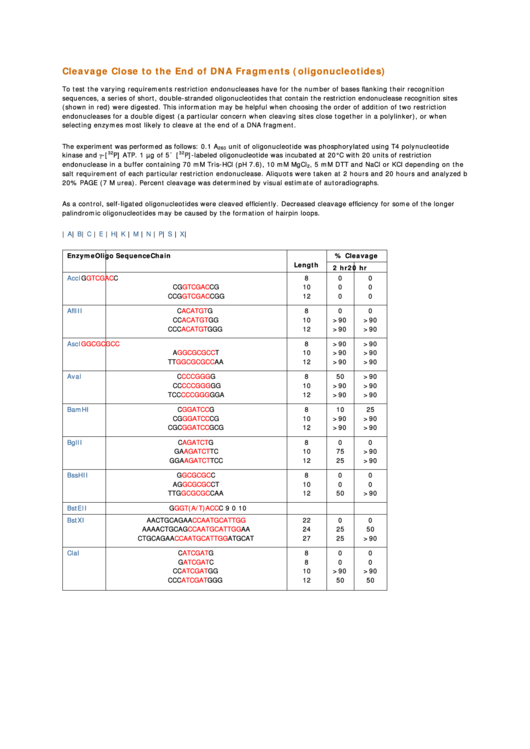
Cleavage Close to the End of DNA Fragments (oligonucleotides)
To test the varying requirements restriction endonucleases have for the number of bases flanking their recognition
sequences, a series of short, double-stranded oligonucleotides that contain the restriction endonuclease recognition sites
(shown in red) were digested. This information may be helpful when choosing the order of addition of two restriction
endonucleases for a double digest (a particular concern when cleaving sites close together in a polylinker), or when
selecting enzymes most likely to cleave at the end of a DNA fragment.
The experiment was performed as follows: 0.1 A
unit of oligonucleotide was phosphorylated using T4 polynucleotide
260
32
32
kinase and γ-[
P] ATP. 1 µg of 5´ [
P]-labeled oligonucleotide was incubated at 20°C with 20 units of restriction
endonuclease in a buffer containing 70 mM Tris-HCl (pH 7.6), 10 mM MgCl
, 5 mM DTT and NaCl or KCl depending on the
2
salt requirement of each particular restriction endonuclease. Aliquots were taken at 2 hours and 20 hours and analyzed by
20% PAGE (7 M urea). Percent cleavage was determined by visual estimate of autoradiographs.
As a control, self-ligated oligonucleotides were cleaved efficiently. Decreased cleavage efficiency for some of the longer
palindromic oligonucleotides may be caused by the formation of hairpin loops.
|
A
|
B
|
C
|
E
|
H
|
K
|
M
|
N
|
P
|
S
|
X
|
Enzyme
Oligo Sequence
Chain
% Cleavage
Length
2 hr
20 hr
AccI
GGTCGACC
8
0
0
CGGTCGACCG
10
0
0
CCGGTCGACCGG
12
0
0
AflIII
CACATGTG
8
0
0
CCACATGTGG
10
>90
>90
CCCACATGTGGG
12
>90
>90
AscI
GGCGCGCC
8
>90
>90
AGGCGCGCCT
10
>90
>90
TTGGCGCGCCAA
12
>90
>90
AvaI
CCCCGGGG
8
50
>90
CCCCCGGGGG
10
>90
>90
TCCCCCGGGGGA
12
>90
>90
BamHI
CGGATCCG
8
10
25
CGGGATCCCG
10
>90
>90
CGCGGATCCGCG
12
>90
>90
BglII
CAGATCTG
8
0
0
GAAGATCTTC
10
75
>90
GGAAGATCTTCC
12
25
>90
BssHII
GGCGCGCC
8
0
0
AGGCGCGCCT
10
0
0
TTGGCGCGCCAA
12
50
>90
BstEII
GGGT(A/T)ACCC
9
0
10
BstXI
AACTGCAGAACCAATGCATTGG
22
0
0
AAAACTGCAGCCAATGCATTGGAA
24
25
50
CTGCAGAACCAATGCATTGGATGCAT
27
25
>90
ClaI
CATCGATG
8
0
0
GATCGATC
8
0
0
CCATCGATGG
10
>90
>90
CCCATCGATGGG
12
50
50
ADVERTISEMENT
0 votes
Related Articles
Related forms
Related Categories
Parent category: Medical
 1
1 2
2 3
3